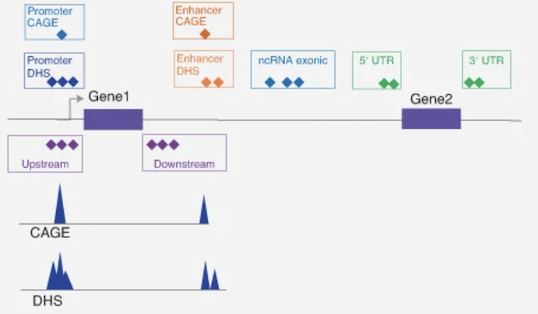
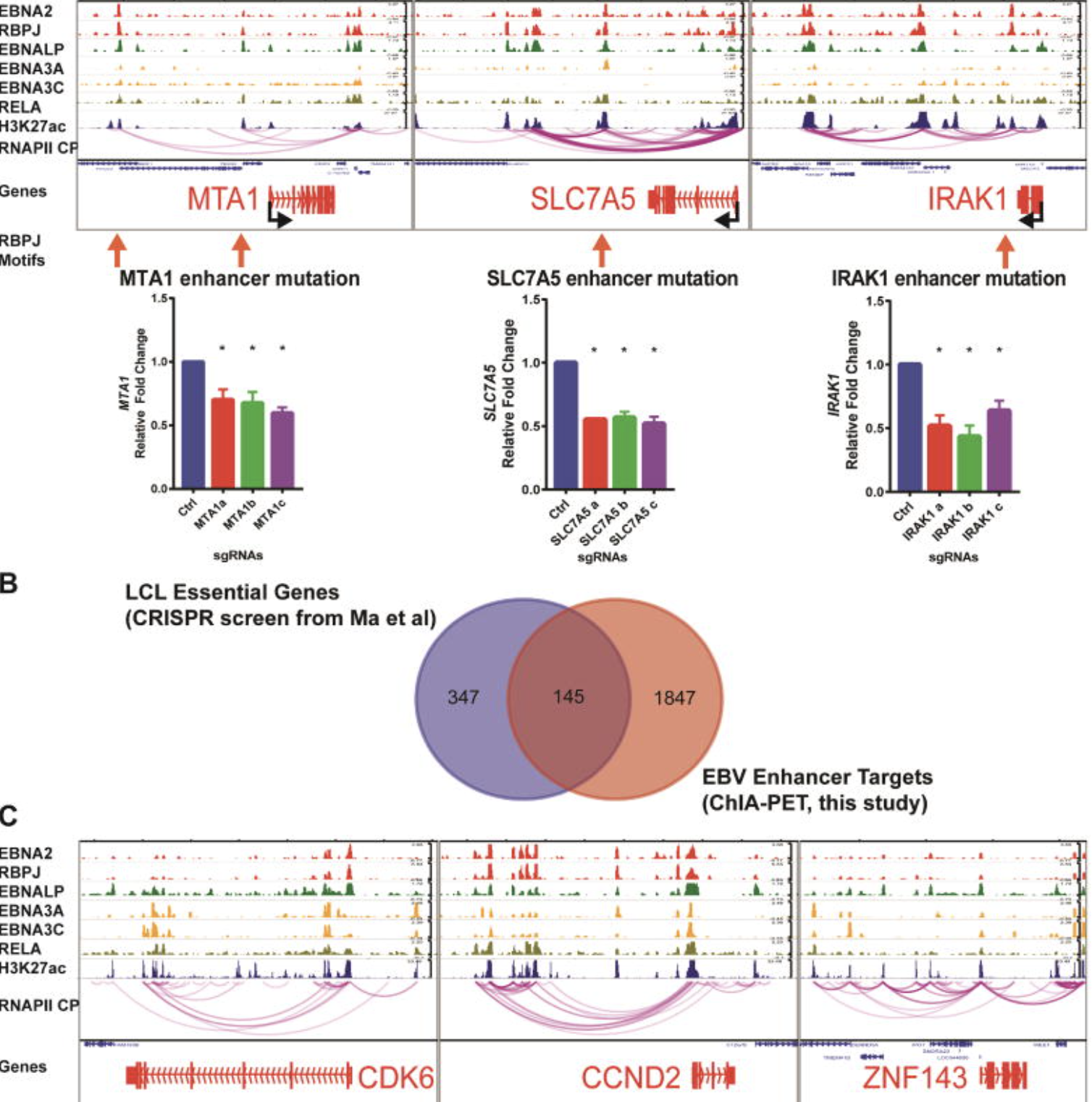
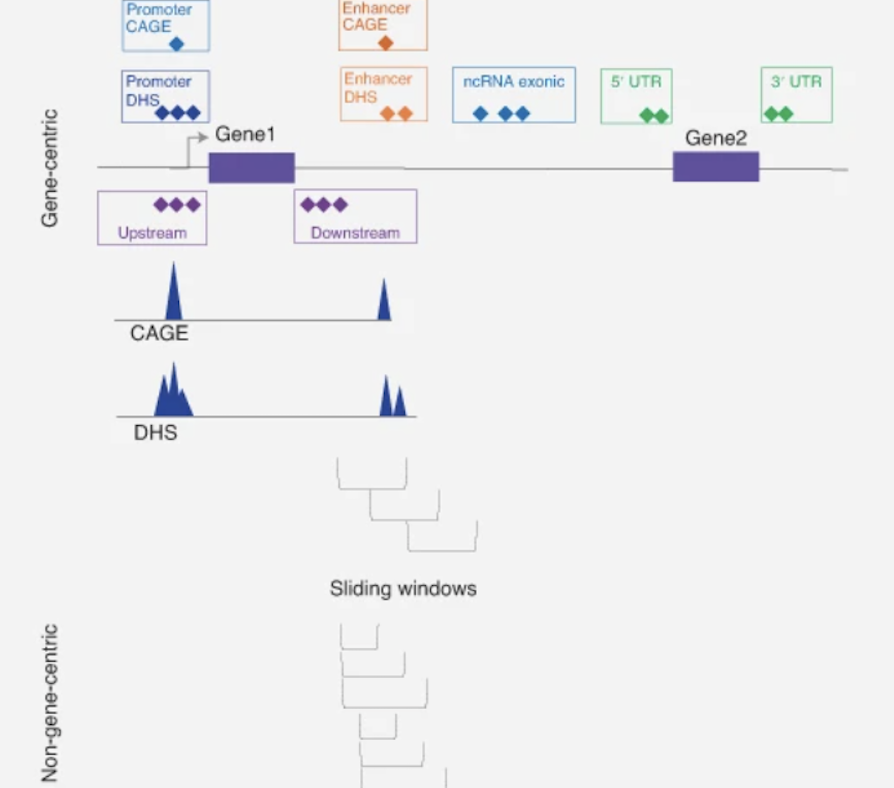
A statistical framework for multi-trait rare variant analysis in large-scale whole-genome sequencing studies.
Li X, Chen H, Selvaraj MS, Van Buren E, Zhou H, Wang Y, Sun R, McCaw ZR, Yu Z, Jiang MZ, DiCorpo D, Gaynor SM, Dey R, Arnett DK, Benjamin EJ, Bis JC, Blangero J, Boerwinkle E, Bowden DW, Brody JA, Cade BE, Carson AP, Carlson JC, Chami N, Chen YI, Curran JE, de Vries PS, Fornage M, Franceschini N, Freedman BI, Gu C, Heard-Costa NL, He J, Hou L, Hung YJ, Irvin MR, Kaplan RC, Kardia SLR, Kelly TN, Konigsberg I, Kooperberg C, Kral BG, Li C, Li Y, Lin H, Liu CT, Loos RJF, Mahaney MC, Martin LW, Mathias RA, Mitchell BD, Montasser ME, Morrison AC, Naseri T, North KE, Palmer ND, Peyser PA, Psaty BM, Redline S, Reiner AP, Rich SS, Sitlani CM, Smith JA, Taylor KD, Tiwari HK, Vasan RS, Viali S, Wang Z, Wessel J, Yanek LR, Yu B, NHLBI TOPMed Consortium, Dupuis J, Meigs JB, Auer PL, Raffield LM, Manning AK, Rice KM, Rotter JI, Peloso GM, Natarajan P, Li Z, Liu Z, Lin X.
Nat Comput Sci. 2025;5(2):125-143.