Machine learning for biomedical data
AI Genomics and Digital Pathology
AI-driven approaches for pathogenic variant annotation, quality control in large-scale WGS, multi-omic integration, and digital pathology risk prediction.
Overview
AI Genomics and Digital Pathology
The CV highlights an expanding translational data-science direction: applying machine learning and deep learning across variant annotation, functional genomics, automated WGS quality control, and whole-slide pathology image analysis. This adds a forward-looking AI thread to the job-market research profile while staying grounded in genomics and disease biology.
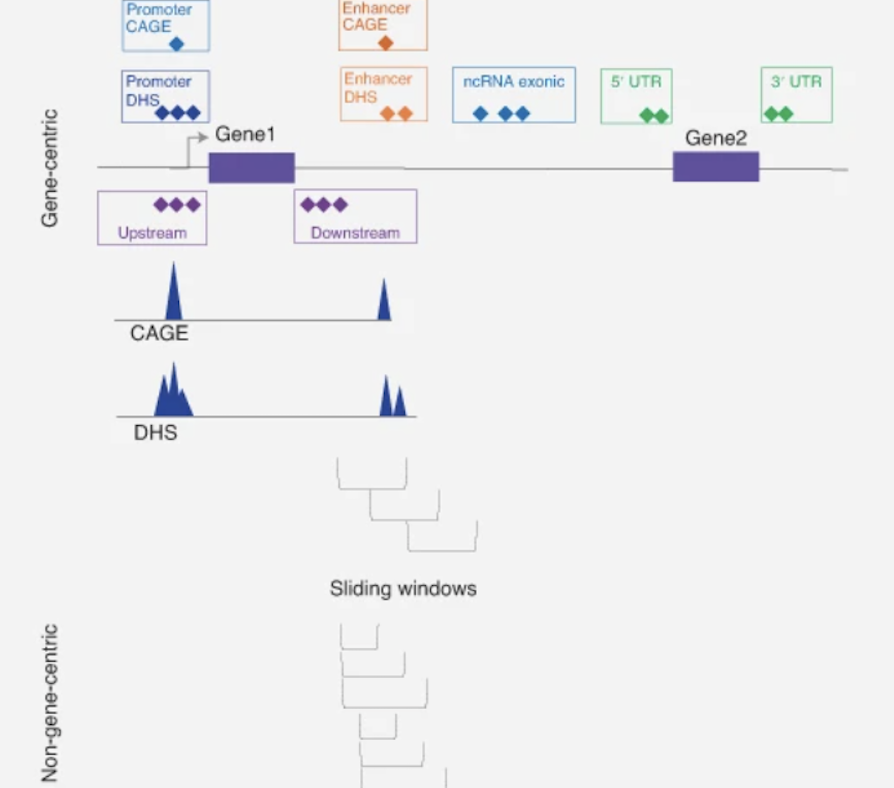
- Pathogenic noncoding variant annotation using statistical and neural-network models that integrate epigenetic marks, conservation, microRNA binding, and other multi-omic data.
- Automated WGS quality-control pipelines with variant-level and sample-level filters, anomaly detection, and batch-effect checks.
- Digital pathology work using H&E whole-slide images for tumor-region localization and 10-year progression-risk stratification in ER-positive, lymph-node-negative breast cancer.
- Graph-based and machine-learning approaches for regulatory-network reconstruction, protein-protein interaction modeling, pathway enrichment, and clinical multi-omic analysis.
Publications
Related work
Representative publications connected to this project.
2024
2024
2022
A multi-dimensional integrative scoring framework for predicting functional variants in the human genome.
Am J Hum Genet. 2022;109(3):446-456.