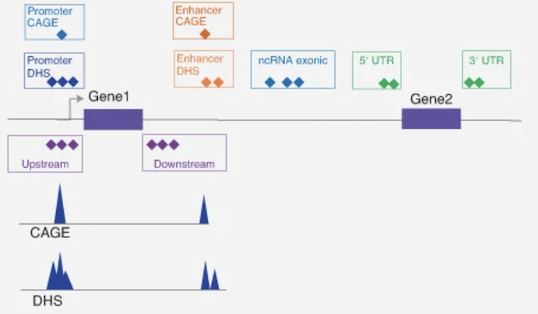
Population Genetics
Research on scalable whole-genome sequencing analysis, rare-variant association testing, functional annotation, and genetic discovery in diverse populations.
Explore projectProjects and research directions
A clearer map of the research program, with each major area connected to publications, software, and project detail pages.
Overview
Each project area links methods, biological questions, representative publications, and software or data resources.

Research on scalable whole-genome sequencing analysis, rare-variant association testing, functional annotation, and genetic discovery in diverse populations.
Explore project
FAVOR is a functional annotation resource and software ecosystem for interpreting human genetic variation across the genome.
Explore project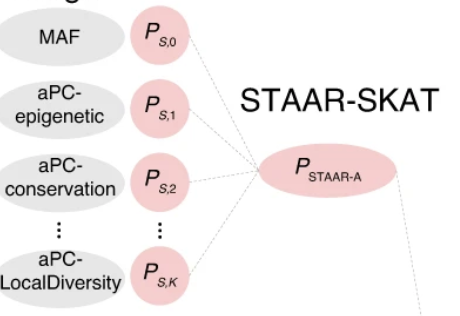
STAAR and its extensions use functional annotation to improve rare-variant association analysis for whole-genome sequencing studies.
Explore project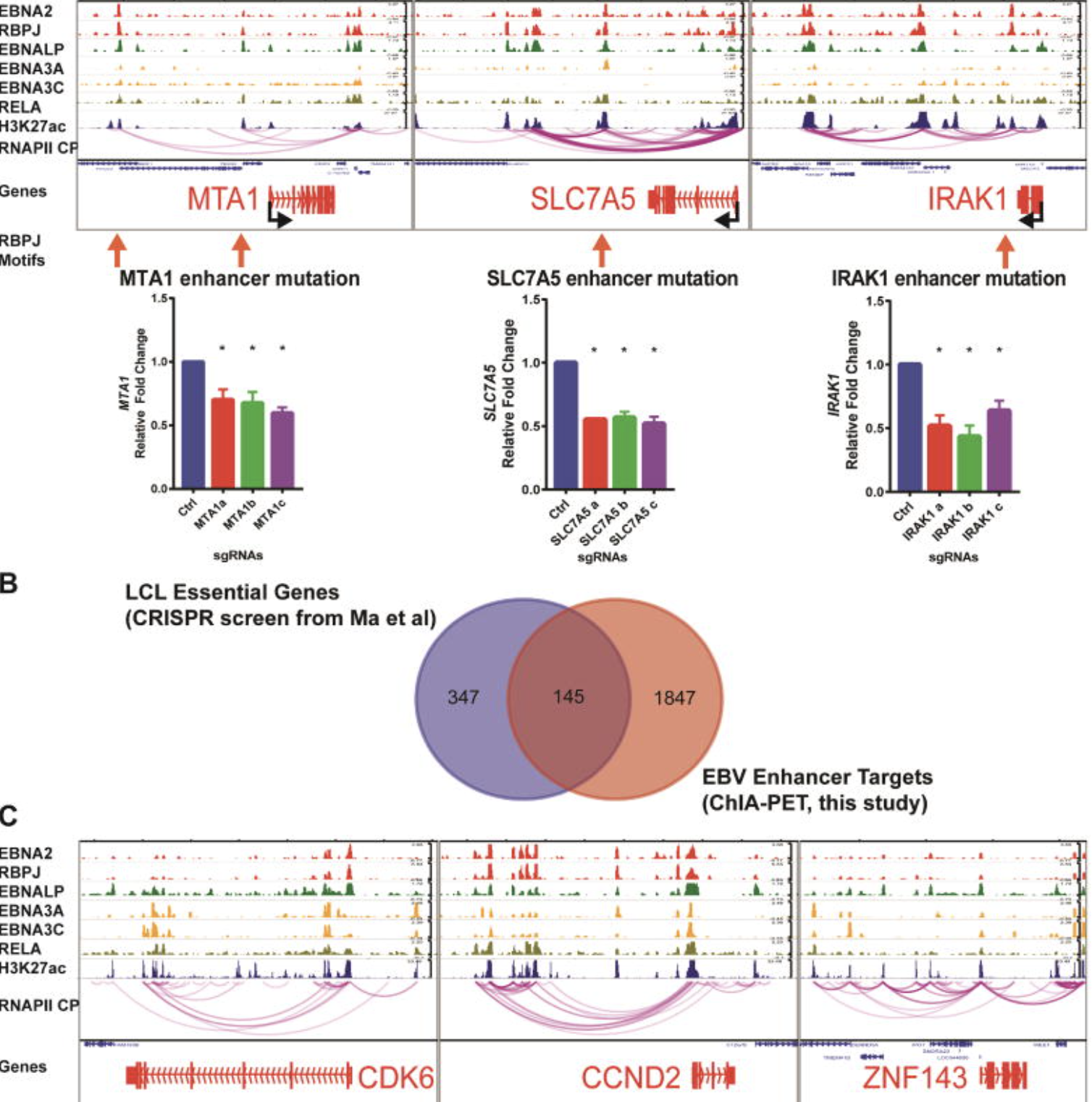
Integrative functional genomics of Epstein-Barr virus regulation, host chromatin, and cancer-relevant transcriptional programs.
Explore project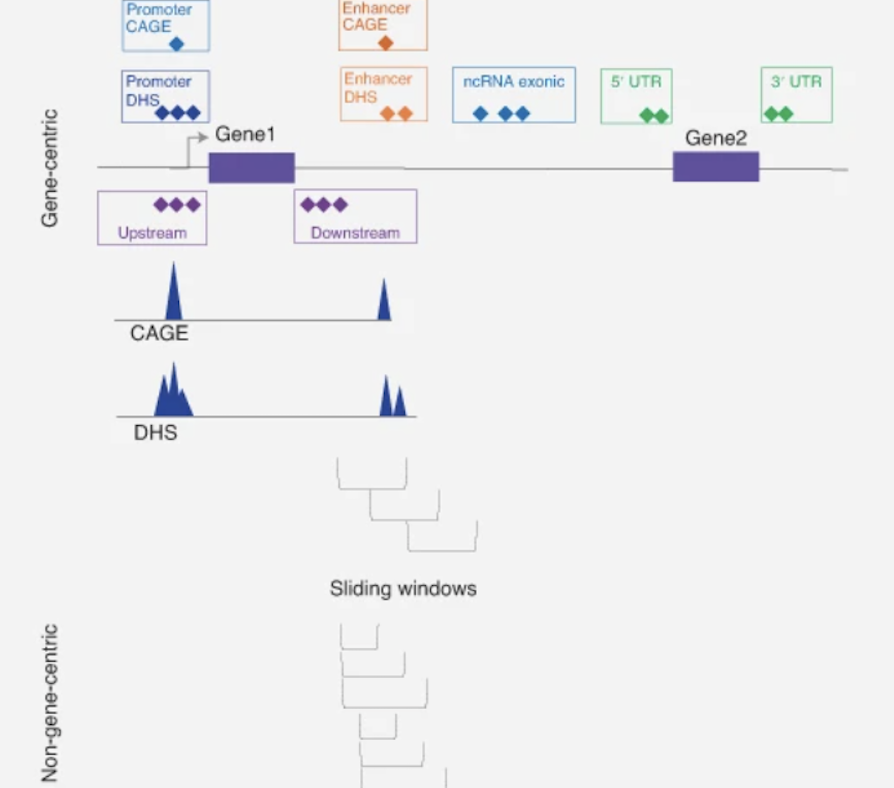
AI-driven approaches for pathogenic variant annotation, quality control in large-scale WGS, multi-omic integration, and digital pathology risk prediction.
Explore project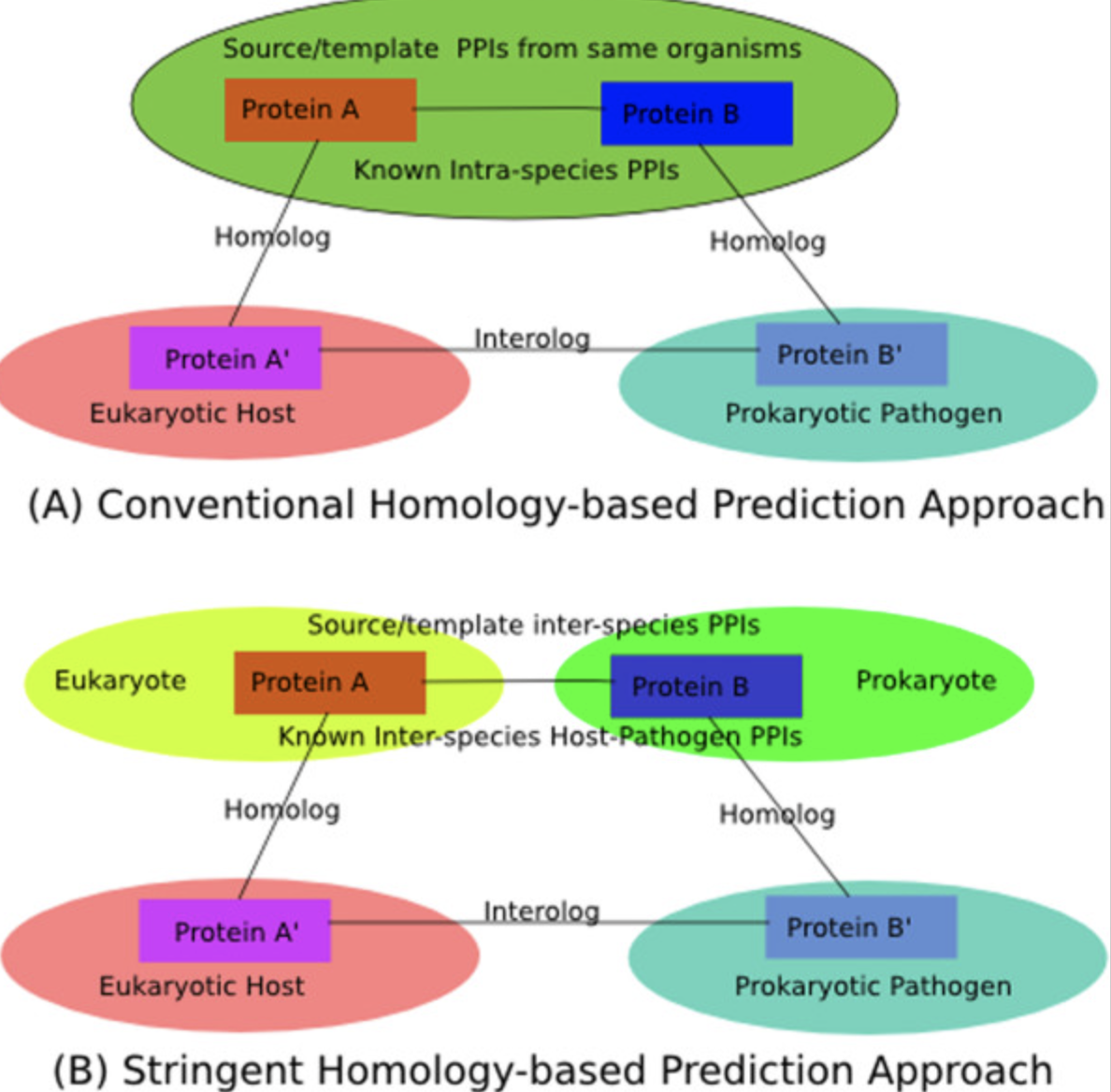
Computational prediction, assessment, and biological interpretation of protein-protein interaction networks.
Explore project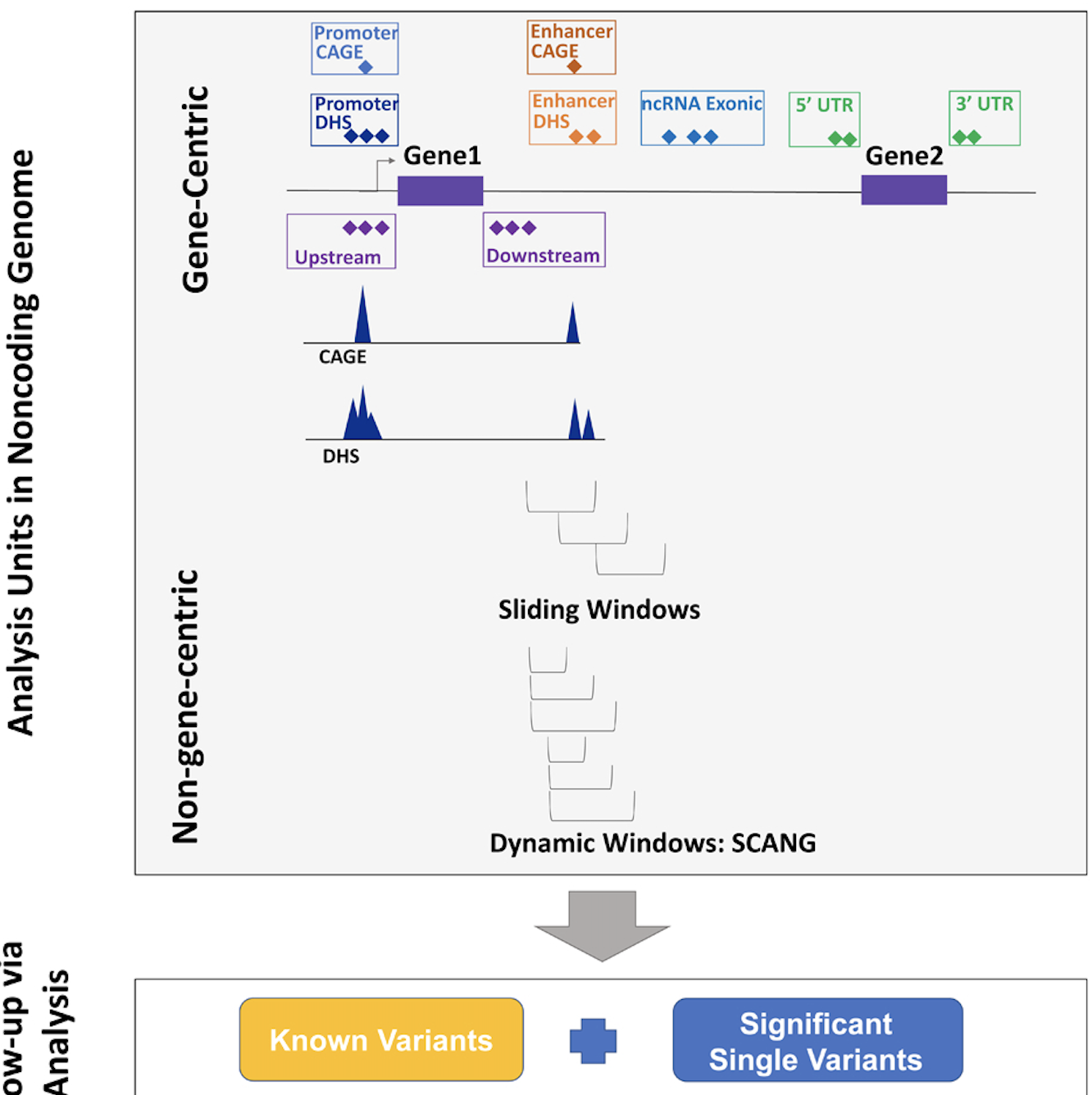
From methods to software
The research program connects method development, statistical inference, functional genomics, scalable software, and public resources. That makes the portfolio legible as both a scientific agenda and an engineering record.